Why is the process of genome duplication so prevalent in plants?
Polyploidy and its ecological consequences
Polyploidy is associated with some of the largest ecological changes resulting from a single genetic event. These changes include dramatic shifts in environmental tolerances, geographic range and ecological interactions. The mechanisms and rate by which polyploidy achieves such changes, and whether there are predictable changes, are less understood.
Our lab has made a number of contributions to this general problem. Through the work of former Ph.D. student, Dr. Sara Martin, we have examined the link between polyploidy and geographic range size and position in a large scale comparative analysis, the genetic basis of shifts in geographic distribution, and compared the ability of diploid and tetraploid populations to respond to selection. More recently, in collaboration with Dr. Hafiz Maherali, we have been studying the physiological consequences of whole genome duplication, in particular on drought tolerance and cell storage capacity.
Researchers: Evan Pacey (Ph. D. candidate), Sara Martin (Ph. D.)
Unreduced (2n) gamete production and polyploid formation
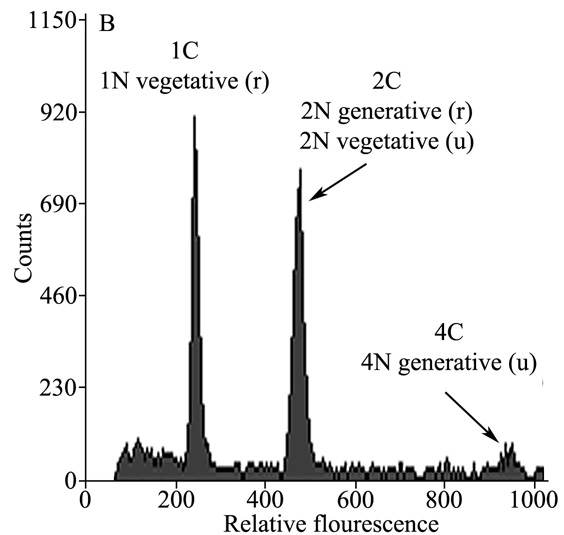
Polyploidy (duplication of genome copies / nucleus) is one of the most common genetic mechanisms of ecological and evolutionary change in plants. Despite its importance, relatively little is known about how these duplications arise in natural populations. We believe polyploids are often formed through the production and union of unreduced (2n) gametes but little is known about the factors controlling this process. Our lab has been tackling this problem from several perspectives. We have been developing new, high throughput, methods for measuring unreduced pollen using flow cytometry, quantifying the magnitude of unreduced pollen production and its sources of variation within and among natural populations, and evaluating the role of genetic and environmental sources of variation in 2n pollen production in controlled environments.
Researchers: Paul Kron (Research Assoc.), Julia Kreiner (UG), Dylan Sora (UG)
Polyploidy evolution and speciation
Polyploid evolution is associated with ~15% of all plant speciation events; however, the population process by which polyploids establish, and the rate (abrupt or gradual) and mechanisms by which reproductive isolation evolves are rarely studied.
Over the last 15 years, we have investigated the obstacles to having rare polyploids establish in the presence of their diploid progenitors, and explored a wide range of attributes (assortative mating, polyploidy fitness) that enable polyploids to overcome these obstacles. In particular, we have examined the contributions of flowering phenology, plant-pollinator interactions, competitive ability, self-fertilization, and gametic competition to polyploidy success. Most recently, Wendy Van Drunen is examining how the extent and mode of clonal reproduction can influence polyploidy establishment and subsequent evolutionary modification. A unique element of our research is the use of synthetic polyploids as well as naturally occurring ones to understand the direct impacts of whole genome duplication on polyploidy speciation.
Researchers: Wendy Van Drunen (Ph. D. candidate), Ken Thompson (UG), Tracy Burton (Ph. D.), Holly Sabara (Ph. D.), Sarah Baldwin (M. Sc.), Yvette Roy (M. Sc.)
Polyploidy and its genetic / genomic consequences
Polyploidy-mediated adaptation and speciation can not be fully understood using ecological and phenotypic approaches alone. We have been using genetic approaches to address two questions: 1) what are the genetic relationships between diploids and polyploids in the same species, and what can that tell us about the number and location of polyploid origins, and the potential for gene flow among ploidy levels: and 2) what is the genetic basis of phenotypic change in polyploids? Does gene expression change in proportion to the number of copies (gene dosage)? How does genotype identity modify the impact of whole genome duplication on phenotypic traits.
Genetic research in our lab is just beginning to develop. Thus far, former M.Sc. student, Yvette Roy, and postdoctoral fellow, Justin Ramsey, examined the population genetics and relationships among diploid, triploid and tetraploid fireweed (Chamerion angustifolium) and tetraploid and hexaploid yarrow (Achillea millefolium), respectively. Recently, postdoctoral fellow Adam Green developed a study system of 100 diploid accessions and their corresponding synthetic tetraploids of Arabidopsis thaliana for the purpose of examining whether the effects of duplication are consistent or whether they are genotype dependent. Currently, M.Sc. student Werdah Iqbal (under co-supervision of Dr. Joe Colasanti) is building on that study system to determine how gene expression for key flowering time genes changes across different diploid and synthetic tetraploids pairs.
Researchers: Werdah Iqbal (M. Sc. candidate), Adam Green (PDF), Justin Ramsey (PDF) Yvette Roy (M. Sc.)